. Leica SP8 confocal microscope
Leica SP8 Confocal Microscope (owned by ACCM, open access)
Location: Ross 9th Floor
The Leica SP8 confocal microscope is on a Leica DMi8 inverted microscope stand (DMi8CEL).
* open access to JHU (and outside) researchers - if heavily booked, ACCM staff will have priority access (i.e. we want ACCM staff to be able to use at least 20 of the ~40 hour work week hours).
Quick summary:
* Reservations and billing on iLab, $27/hr work hours, $20/hr night and weekends. We bill in half hour increments. Once you know what you are doing, you could acquire "over lunch" (or over long lab meeting), or overnight (or over weekend), unattended -- and use your time better that confocal sitting if you do not need to be one hand.
* Four laser lines: 405, 488, 552, 638 nm.
* Two excellent photon counting detectors, HyD1 and HyD3 (second generation hybrid detectors). Maximum photon counting rate: 60 photons per microsecond - see 2024089F text (URL below) for details and source (search on that date). Data reporting in photon counts are is far superior to standard confocal PMT(s) which report in 'arbitrary units" and whose output values depend on HV (high voltage) and offset -- which are subject to abuse by users.
20240809F Leica SP8 HyD detectors linear range up to 60 photons per second (each) - see http://confocal.jhu.edu/mctips/leica-sp8-hyds-linear-range-to-60-photons-per-microsecond
* Typical immunofluorescence experiment: DAPI, Alexa Fluor 488, Alexa Fluor 568 (or Alexa Fluor 555), Alexa Fluor 647. AF647 is near infrared - not visible by eye, so if you only have DAPI and two immunomarkers, just use Alexa Fluor 488 and Alexa Fluor 568 (or AF555). I note that Alexa Fluor PLUS secondary antibodies from ThermoFisher are 3x brighter, though 1.3x more expensive than plain AF antibodies. To me 3x brighter is a win, since you might have $3 vs $4 in antibodies vs $27/hr confocal time.
* I strongly recommend usingthe HyD detectors (not the "PMT 2") for all fluorescence acquisitions. The non-PC mode has a "Gain" setting, whose default is 10. this is simply the photon count time 10. However, the user can set the gain to some other value, which if not done consistently will make data interpretation more difficult. I added at the bottom of this page a multi-part Confocal listserv thread about HyD Detectors (February 2024).
|
Leica SP8 Confocal Microscope 20250314 Update - Users should read. * I updated the PC to Windows 11. * Replaced "K:" 3 GB/sec data drive (~3 TB) with new "D:" 7 GB/sec data drive (4TB). Also added a "Leica Temp drive" -- for Leica software to use, not users - also 7 GB/sec (4TB) (crystaldiskmark benchmark software, not necessarily open/save/transfer to JH OneDrive / transfer to Research I.T. SafeStor). the Highpoint SSD7105 NVMe drive array could have been configured for 14 GB/sec (speed of PCIe gen3 x16 slot) - our Leica rep explained to me that better read/write performance by splitting. * An unexpected glitch is that the Process menu "Mosaic Merge" for stitching "tile scan" datasets no longer works ("hangs" the PC - requiring PC restart). Also hangs if turn on Auto-Stitch in tile Scan. Our local leica reps, Kevin and Peter, worked through options and found that: Note: use 91.84 degree rotation (matches eyepieces and motorized stage), leave Swap and Flip options off for NAVIGATOR-Mosaic Merge. 1. Acquire using HyVolution2 or TCS SP8 mode tile scan (leave off auto-stitch) (HyVolution2 better because you can load your settings and because Fast Live button). --> after acquisition, always save your LIF file!!! 2. Switch to TCS SP8 mode.
--> after "MM", always save your LIF file!!! If the LIF file is big (1+ GB may overwhelm your lab/office PC), I suggest you save one or few tile scans and stitched data,
Tip: If using HyVolution2 GPU deconvolution (3D image acquisition), you are most likely better off acquiring and GPU deconvolving many modest size tiles, say 4x4 tiles of 1500x1500 pixels each (or 8x8 tiles of 750x750 pixels each), than one giant low zoom 6000x6000 pixels. This is because our NVidia GeForce M6000 24GB GPU has a good amount of ram, but not massive amount of RAM. The deconvolution will be more efficient with many small tiles. I also note we default to 10% overlap, so two adjacent 1500x1500 tiles will result in an ~2850x1500 image. FYI – someday have Kevin Murphy (Leica confocal sales) or I show you how to acquire in NAVIGATOR (the standard tile scan is a much simpler interface). Kevin and I will probably revisit the ’standard but not working mosaic merge’ issue in a few weeks. No promises we will get it to work—might be a Windows 11 incompatibility that may be unsolvable. I am happy Kevin and his colleague Peter found NAVIGATOR’s Mosaic Merge works - per above. |
|
HyVolution mode (turn off the "red chain" button to enable changing various settings. 600 Hz Format (#pixels): whatever you need Zoom (scanner zoom): whatever you need. Format * zoom combine for pixel size, 20x 0.75NA ... 120nm 63x 1.35NA ... 60 or 50nm (50 nm if follow-up with deconvolution). Averaging: 1. Line accumulation: start with 10, do more if dim signal (of positive control), less if very bright, ideal 300 or more counts per pixel.
|
June 1, 2018 was the start of the iLabs Organizer reservations for our Leica SP8 confocal microscope (much more efficient than the old reservation and billing system):
ACCM Leica SP8 Confocal Microscope (ACCM Confocal Microscope)
https://johnshopkins.corefacilities.org/equipment/345585/?tab=schedule#
If a user needs their PI or administrator to set up account numbers (IO#'s in JHU accounting jargon), see
https://help.ilab.agilent.com/36900-managing-your-group/279959-membership-requests-fund-numbers

This microscope is owned by the Department of Anesthesiology & Critical Care Medicine (ACCM) and managed by Ross Fluorescence Imaging Center, - Director Prof. Bin Wu, image core manager George McNamara, PhD, 305-764-2081 (cell), gmcnama2@jhmi.edu.
Suggested text for Acknowledgements:
Microscope: We acknowledge the Department of Anesthesiology & Critical Care Medicine (ACCM) for access to their Leica SP8 confocal microscope.
If you get 'significant' assistance mage core staff: We thank George McNamara, Ph.D. for assistance with the ACCM confocal microscope (routine training and question answering does not rate acknowledgement).
Our Lecia SP8 confocal microscope was purchased on a budget, so between the price and capabilities of a 'baby confocal' like Leica SPE, and a 'fully equipped' confocal microscope (example: our SP8 has only 3 detectors, no incubator ... with 3 detectors, can acquire many channels by sequential scanning multiple tracks). The two Leica HyD detectors are best operated in "counting mode" = photon counting. This enables much simpler quantitation of fluorescence signals than conventional PMTs whose digitizers output values that depend on both the High Voltage ("HV") gain and the "offset".
- Our SP8 has space for two more HyD's internally. Our HyD's are "2nd generation", in mid-2018 Leica introduced single molecule detection SMD HyD's, that each count 2x faster than our 2nd gen HyD's. If you have money, and especially if you need speed and/or plex, please consider purchasing SMD HyD's for this SP8 confocal microscope. Ideally purchase FOUR SMD HyD's and negotiate trade-in of the current two 2nd gen HyD's. This would enable "fast photon counting" with the four SMD HyD's (8x faster counting than now). I note that SMD HyD's are also FLIM (fluorescence lifetime imaging microscopy) capable, so if a FLIM compliant pulsed laser (ex. 440 nm 80 MHz, or fiber laser) was purchased, could do "fast FLIM" (would also need upgrade of LAS X software to LAS FALCON fluorescence lifetime contrast). We have the newer Klondike scanner with linear scanning electronics, not the older sinusoidal "Fermi" FOV scanner electronics (info on scanner electronics from Steffen Dietzel post on confocal listserv on 20220221Mon.
The Department of Anesthesiology & Critical Care Medicine (ACCM) users are aiming to use 20 hours per week, that is, 50% of JHU business hours access (it's ok if ACCM uses more). It is a 'win' for everyone if we can maximize efficient use of the SP8 since more revenue helps pay for the instrument and service contract. For now, we will try to manage this ACCM 'preferred access' informally. (i) If ACCM users get above 20 hours per work week: great. (ii) if non-ACCM users add up to more than 20 hours in any work week and ACCM users did not need 20 hours: great for everyone. (iii) We encourage ACCM users to reserve well in advance (i.e.1+ week).
This is a state of the science/engineering confocal microscope - we want everyone who can benefit from the LeicaSP8 confocal microscope to be able to get excellent data with it.
Rates all users are:
* $27/hour for fully trained expert users, during JHMI business hours (Mon-Fri 9am-5pm).
. * $20/hour for fully trained expert users, outside JHMI business hours (outside Mon-Fri 9am-5pm).
* $70/hr for users, with Dr. McNamara's oversight.
* $120/hr for Dr. McNamara to provide training and/or to operate the confocal microscope for users who need an operator.
==> Dr. McNamara reviews sessions on the sign-in sheet when doing the 'confirm' step in iLab Organizer.
Common 2*2 immunofluorescence experiment:
scan track 1: 488 nm and 638 nm lasers, AF488 on HyD1, AF647 on HyD3.
scan track 2: 405 nm and 552 nm lasers, DAPI on HyD1, AF568 on HyD3.
HyD's in photon Counting mode, usually 10 line accumulation, no frame accumulation. If dim signal: 16 line accumulation and 'whatever needed' frame accumulation you need to get good data (overnight or over weekend if necessary ... though you should think about better fluorophores, more laser power, tyramide signal amplification, etc, to get better signal).
widefield microscope resolution equation: dxy = 0.61 * Lambda / NA ... Nyquist theorem (2D) suggests ~3x sampling for pixel size.
confocal microscope resolution equation (1.0 Airy unit standard of care); dxy = 0.51 * Lambda / NA ... ... Nyquist theorem (2D) suggests ~3x sampling for pixel size.
|
Objevtive lens Confocal zoom range 0.75 - 48.0, so 63x lens is effective 42x - 3024x (not that anyone will benefit from latter zoom). |
XY Pixel | Z-Step |
Working distance |
comment |
| 5x / 0.15 | 600 nm | 1800 nm | 12.0 mm | large field of view |
| 20x / 0.75 | 120 nm | 360 nm | 0.62 mm = 620 um | standard dry objective lens |
| 63x / 1.40 NA (oil immersion) | 50 nm | 150 nm | 0.14 mm = 140 um | oil immersion objective lens |
|
Our 10x lens was not purchased for confocal microscopy so we choose to not provde recommendations for this lens. |
||||
|
Below: SVI.nl recommendations for the 63x/1.4NA lens for Brilliant Violet 421: 405nm excitation, 440 nm emission Smaller pinhole sizes expected to reduce light level -- by how much is NOT simply proprotional to relative pinhole area: best to measure for yourself. For highly photostable fluorophores (or reflected light mode), could "go small". Potentialy stable fluorophores include BV421 (above), Phitonex NovaFluors (ThermoFisher), StreamBioUK CPNs. |
Relative XY resolution improvement (approximate - Zeiss has a nice graph on this in a PDF brochure) | |||
|
63x / 1.40 NA (oil immersion) pinhole 1.0 Airy Unit |
37 nm | 112 nm | 1 | |
|
63x / 1.40 NA (oil immersion) pinhole 0.666 Airy Unit * Jeff Reese, NIDDK / NIH, wrote in Confocal Listserv wrote they like "0.6 - 0.7 Airy unit as sweet spot" for confocal. I simplify here to "sweetest spot of 0.666 Airy unit. |
36 nm | 108 nm | 1.05 | |
|
63x / 1.40 NA (oil immersion) pinhole 0.5 Airy Unit |
36 nm | 108 nm | 1.10 | |
|
63x / 1.40 NA (oil immersion) pinhole 0.3 Airy Unit |
36 nm | 108 nm | 1.20 |
** See also https://svi.nl/NyquistCalculator for SVI (Huygens) recommendations for optimal XY pixel and Z-step size. Enable "Calculate a Point Spread Function" to have the web page compute and display XY and XZ images of point spread. GM recommends specimen refractive index 1.515 for oil immersion objective lenses (both in calculation and as your mounting medium). We have "HyVolution2" = Leica SP8 confocal and Huygens Essential. See also their "Explanation: proper images" and additional text on their web page.
For 20x/0.75NA objective lens: 1 Airy unit pinhole size, 120 nm XY pixel size, 360 nm Z-step size (GM's simple XY has three times better spatial resolution than Z).
For 63x/1.4NA objective lens -- with HyVolution2 set to 'aggressive', improvement is for each axis, so 1.2^3 = 1.728 or 1.414^ = 2.82, for volume):
"standard confocal" resolution: 1.0 Airy unit pinhole size, 50 nm XY pixel size, 150 nm Z-step size.
"super-resolution" resolution (~1.2x improvement): 0.5 Airy unit pinhole size, 40 nm XY pixel size, 120 nm Z-step size. Note: 0.5 AU and "Area = pi r^2" implies 25% intensity.
"super-duper resolution" resolution (1.414x improvement): 0.3 Airy unit pinhole size, 40 nm XY pixel size, 120 nm Z-step size. Note: 0.3 AU and "Area = pi r^2" implies 9% intensity.
Reminder: the HyD detectors photon counting mode greatly facilitates quantitation of these settings. You can evaluate each by enabling sequential scan tracks with each pinhole setting.
Equipment Summary:
Scanner speed 1 to 1800 Hz, default 400 Hz, typically used at 600 Hz to enable full field imaging (zoom 0.75x). Hz = scan lines per second.
Faster speeds require zooming. For example, 1800 Hz requires zoom 7.5x or more (you control number of pixels). We encourage evaluating using "max speed" 1800 Hz with summing the HyD photon counts (line accumulation = 10, for example). Max speed decreases photobleaching (speed and scanning area interact).
Hardware zoom 0.75x - 48x (by changing confocal scanning area with the XY scanning mirrors).
Max image resolution 8192 x 8192 pixels (64 megapixels). Max number of scan tracks is ~40 in LAS X (9/2018 version). Since we have four laser lines, generally two, three or four scan tracks is sufficient. There is an channel unmixing "matrix" to let you unmix 'spectral overlaps'.
4 lasers (power out of laser, when new - some power settings "after objective lens" in box below):
405 nm (50 mW)
488 nm (20 mW)
552 nm (20 mW)
638 nm (30 mW)
Note: HyD's detection must be at least 5 nm from laser lines, for simplicity, for SAFETY please restrict your HyD emission bands to at least 10 nm from any laser line. Example, 500-540 nm is away from 488 and 552 nm laser lines.
|
We typically operate the 405nm laser at 0.5% or 1.0% laser power and the 488, 552, 638nm lasers at 1%, 2%, 3%, 4% or 5% (most often 1 or 2%). Higher power settings may result in photobleaching the field of view. To demonstrate photobleaching, GM typically: 1. at 1x zoom, 1% laser power, usuallyHyVolution "fast live" (512x512 pixels, no line accumulation), positions a brightly fluorescent cell "dead center" in the field of view. 2. Zoom to 5x and re-center cell if needed. 3. Increase laser power to 10% or 25% or 50% (if HyD detector safety interlock triggers, top stanning, decrease laser power, start scanning again). 4. Increase zoom to 20 or 40 or 48 (max is 48), and watch the fluorescence signal decrease (yes, could operate in time acquisition mode - for simple proof of photobleaching no need). 5. "Just look" as signal 'just fades away'. 6. Decrease zoom. GM routinely suggests to users they could sign - or faster just write initials - into cell(s) or tissue - using zoom, laser power and stage to photobleach (simplest if their initials are II or even simpler if they just go by I). ** laser power 09/26/2023 Previous 05/04/2022 |
TCS LIAchroic beamsplitters
3 epi-illumination detectors: 2 HyD, one conventional PMT ... used for fluorescence. Tghe PMT can be used for reflected light (the HyD's need to be protected from reflected light).
Two HyD's = hybrid GaAsP faceplate PMT front end, Aavalanche photodiode (APD) back end), 2x quantum efficiency compared to standard PMTs.
- ==> The HyD's are photon counting devices. This provides a straightforward path for intensity quantitation compared to conventional PMTs.
- We typically recommend 10 line acumulation, resulting in 'typical' pixel (voxel) counts of 5 to 100 vs "empty" pixel dounts of zero or one or two. A photon count of 10 looks small, but compared to 'background' of zero is quite high. With a standard PMT, it is very difficult to 'translate' the intensity value -- the value depends on the detector gain and offset values (each can be chosen poorly, re https://www.youtube.com/watch?v=VA7J0KkanzM ) -- obtained to either (i) photon counts, (ii) number of fluorophores, or (iii) number of antibody molecules (immunofluorescence) or oligo probes (single molecule RNA FISH.
- On our SP8, you can sit with Dr. McNamara (and/or Leica representatives) and evaluate 'on image core time' the HyD's against each other (HyD1 vs HyD3) and our standard PMT ("PMT2") with different settings (sequential track mode makes this easy).
One PMT (Hamamatsu R9624) with low dark current, 40 MHz sampling of ADC digitizer -- in practice is used by Leica field service engineer(s) for calibration during service visits (ex. annual preventive maintenance visits).
1 transmitted light detector (brightfield, DIC).
Motorized XY scanning stage (control by remotes or software, do not touch the yellow motors).
Z-motor.
HyVolution2 quantitative deconvolution software using NVidia M6000 GPU card ("NEAR instant gratification" - some day we would like to upgrade to NVidia RTX 3080 or 3090 GPU, and new PCIe gen4 PC ... if you have lots of money, please cotnribute to our core). "Super-resolution" possible when using the combination of:
small pinhole (<1.0 Airy Unit ... usually 0.5 Airy unit, can 'go down to' 0.28 Airy unit, signal is proportional to area, so 0.5 Airy unit implies 0.25 photons detected per scan, and 0.3 A.U. is 0.09 photons detected, if other parameters constant)
HyD detector(s): photon counting mode please! 16-bit data mode (HyVolution2 ... Configure ... settings ... 16-bit ... if you leave the default 8-bit, the highest value is 255, which is modest intensity).
Huygens Essential deconvolution (SVI Hyugens Essential intregrated into Leica LAS X software). Uusually use "Huygens Essential Automatic" within the HyVolution2 module of LAS X.
Also possible to deconvolve in Huygens Essential. I usually use "Lightning" icon, standard or "aggressive" (aka high resolution), Save As TIFF, and then use "Linked scales". You can save in HDF5 (32-bit loating point values), which Fiji ImageJ can open. I note that MetaMorph can open 16-bit TIFF (and can open an image directly from LAS X's .LIF container file) but MetaMorph does not read HDF5 and does not deal with 32-bit data.
Combination is rated for 140 nm XY resolution (acquire with 50nm or 40nm pixel size, typically Z = 3 * XY pixel size, so 150 or 120nm), compared to "conventional" confocal ~210 nm XY resolution (at 500 nm emission wavelength ... 405 nm excitation of any of BUV395, BV421, SB436 may enable better resolution).
Widefield XY resolution: dxy = 0.61 * Lambda / NA ... 0.61 * 500nm / 1.4 = 218nm ... I recommend pixel size approximately 218/3 = 73nm ... I would use 60nm or 50nm (on FISHscope we use 108nm to get more photons per pixel).
Confocal 1.0 Airy Unit at 500nm fluorescence emission: dxy = 0.51 * Lambda / NA = 0.51 * 500 nm / 1.3 = 182 nm. ... example: AausFP1 (5x brighter than EGFP), Alexa Fluor 488.
Confocal 1.0 Airy Unit at 430nm fluorescence emission: dxy = 0.51 * Lambda / NA = 0.51 * 430 nm / 1.3 = 157 nm ... Brilliant Violet 421 = BV421, also the "421 component" of the many BV tandems (2 channels).
Confocal sweetest spot: Tom Reece (NIDDK, NIH) on confocal listserv wrote that he often uses pinhole in range of 0.6 to 0.7 Airy unit, he calles "confocal sweet spot" (really confocal sweet range) to get better resolution (by a modest amount) and not much light loss (that ios, if one used infinitely small pinhole, would mathematically and geometrical optically get better resolution, but in practice, infinitely small number of photons, even if image infinity time). I simplify to confocal sweetest spot 0.66 Airy unit.
Confocal 0.66 Airy Unit at 430nm fluorescence emission: dxy = 0.94* 0.51 * Lambda / NA = 0.51 * 430 nm / 1.3 = 148 nm ... that is, I estimate ~6% improvement in resolution at 0.66 Airy unit. Te Leica SP8 makes it easy to create sequential scan tracks with different pinhole size, so you can test it yourself.
Deconvolution -- i.e. Huygens HyVolution2 (see above) -- can improve resolution by ~10%, depending on a lot of things, including perfect specimen preparation (as a user, are you perfect?), especially refractive index matching (1.518 for Leica 63x/1.4 NA objective lens, Leica immmersion oil (R.I. 1.518) and your mounting medium ... ThermoFisher Prolong Glass, after correct hardening process, comes very close ... you could also use "TDE" 2,2'-thiodiethanol, Staudt ... Hel 2006 MRT).
Confocal --> Deconvolution 0.66 Airy Unit at 430nm fluorescence emission: dxy = 0.9* 0.94* 0.51 * Lambda / NA = 0.51 * 430 nm / 1.3 = 133 nm.
In practice; good luck getting anywhee close to that resolution on biological specimens ... and in a building that vibrates (all buildings do). If you need super-resolution (beyonf standard of care confocal 1.0 or 0.66 Airy unit), consider using one of the other microscope facilities on campus -- i.e. MicFac -- that have found money for super-resolution. You can also look into DNA-PAINT and using our FISHscope for single molecule localization microscopy.
Confocal quality objectives: HC PL APO 20x/0.75 CS2 and HC PL APO 63x/1.40 Oil CS2 We also moved a pair of Leica 'non-confocal' objective lenses onto the microscope to facilitate 'by eye' searching.
| Objective lens | Magnification |
numerical Aperture (NA) |
Working Distance (WD) |
Confocal quality |
Comment see above for recommended pixel and Z-step sizes for each objective lens |
| HC PL FLUOTAR | 5x | 0.15 | 12.0 mm | good |
Very nice for low mag confocal imaging. Re: McNamara et al 2014 MMB Low magnification confocal microscopy ... |
| N Plan | 10x | 0.25 | 5.8 mm | not | Repurposed from image core lens collection. Useful; for looking. Yes, you can acquire an ok image. |
| HC PL APO | 20x | 0.75 | 0.62 mm | CS2 | Excellent dry objective lens. |
| HC PL APO oil | 63x | 1.40 | 0.14 mm | CS2 |
Excellent objective lens. SP8 offers 0.75x - 40x zoom, so effectively 42x 'total mag' to 'very high' zoom. Refractive index of the Leica immersion oil is 1.518. |
| GM note: standard coverglass is #1.5, ~170 um thick. If you use #0, ~100 um thick, you can get an additional ~70 um working distance, with only modest loss of image quality from standard coverglass. Probably more important to refractive index match your choice of mounting medium (ex. Prolong Glass) and high NA lens immersion medium ... on this microscope, Leica oil R.I. 1.518. |
LAS X 3D visualization.
Please note:
* the Leica SP8 confocal microscope does not currently have an environmental control incubator system. This simplfies specimen access, whle implying most users will work with fixed specimens on microscope slides or 35 mm imaging dishes. GM recommends for growing cells in Mattek or similar 35 mm imaging dishes (#1.5 or precision 170 um coverglass bottom), using minimal amounts of expensive reagents for labeling (antibodies, single molecule RNA FISH), large volume (2 mL) for inexpensive reagent wash steps.
* Having 3 epi-illumination detectors (2 HyD, one conventional) means maximum 3 fluorescence channels per excitation "scan track". The SP8 does enable sequential scan tracks, so you can image more than "3plex".
* We are very open to labs purchasing -- in exchange for time credit -- additional components for the Leica SP8 microscope. For example, 37 C environmental unit (that works on SP8 / DMi8) and two more PMT detectors to "max out" the SP8 scan head. the Leica "laser box" can accept one more laser line (695 nm, possibly ~350 nm). Long term, the SP8 scanhead does not have an X1 port -- this is upgradable in the field. We would love to have 2, 4 or 8 APDs on the X1 port, to enable simultaneous 13plex (5 internal, 8 on X1) acquisition per laser line. APDs are 70% quantum efficiency in the red and near infrared (~600 - 900 nm emission), which would enable further multiplexing, discussed at:
https://www.linkedin.com/pulse/resolution-blues-meets-21plex-salute-fluorescence-basic-mcnamara
https://www.linkedin.com/pulse/bd-biosciences-listed-tandems-horizon-brilliant-violets-mcnamara
We would especially like to see 13plex acquisition per laser line + multiple laser lines + "joint" spatial deconvolution and spectral unmixing, optioanally spectral phasors, per: Hoppe (2008 and 2016), Valm&Borisy (120plex 2016), Fraser ('spectral phasors' 2017) groups publications on parts of these.
Please note that Z-step size is the physical amount the Z-motor moves the objective lens. That is, if imaging air (R.I. 1.0), physical step and "optical step" in the specimen are the same. If you image through a 400 um thick object object that is R.I. 1.600, the 'apparent Z' is 250 um ... that is, the Z-motor will travel 250 um. Example: hemacytometer coverglass is 400 um thick (we measured the thickness 'edge on' by standing the coverglass on edge ... 'mind the gap' ... the hemacytometer counting gap height is 100 um).
***
Leica SP8 scanhead schematic - We have LIAchroic (not AOBS) and two 2nd generation HyD detectors (we have rotation, we do not currently have X1 port). We would love to have a full set four internal SMD HyD detectors (each 2x faster than our 2nd gen, and lower noise because of chiller) and FALCON software upgrade to be able to add multiple detector outputs ... could enable 8x faster imaging ("fast photon counting"), and in future could add AOBS and pulsed laser (ex. Leica "WLL" white light laser [super-continuum laser, 80 MHZ = 12.5 ns interval], to enable precised excitation wavelengths (ex. Nathan Shaner's 6/2019 bioRxiv preprint on AausFP1 bright new GFP could be excited precisely at excitation maximum, AOBS emission side precisely cover emission peak ... and same for the YFP varient).
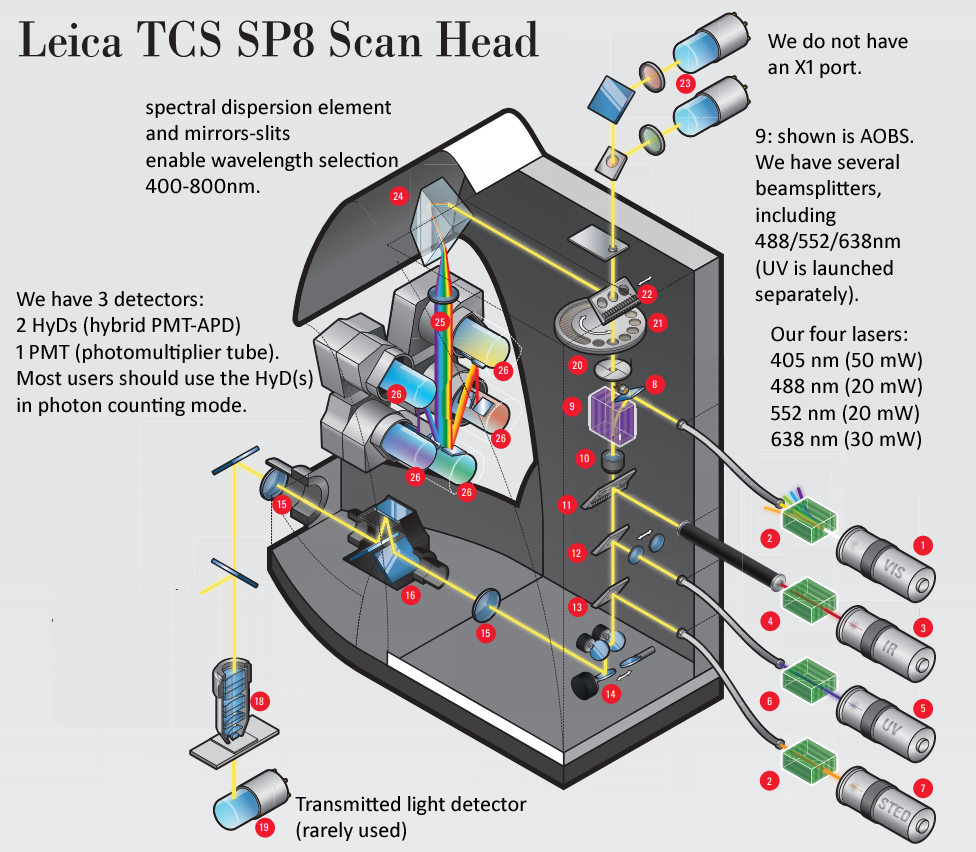
***
News
|
October-November 2020 We hosted Leica STELLARIS confocal microscope demo with: * new "S" detectors (SiPM), ~55% QE (max) and good performance out to NIR. * conventional CW lasers. * Liachroic beamsplitters (similar to our SP8). the model demo'd has some different capabilities than the two featured STELLARIS models (all 3 can have up to 5 S detectors ... or a ix of S and/or HyD 3rd generation visible GaAs or NIR :GaAsP detectors) (note: STELLARIS uses an S detector for field service engineer calibrations, so no needfor any PMT re: SP8). The S detectors and HyD are photon countign and 'fast FLIM' capable. * STELLARIS 5 ... CW and / or pulsed lasers, optional white light laser (WLL) laser (~480 - ~700nm, see data sheet) and AOBS, TauSense "fast FLIM" (fluorescence lifetime imaging microscopy). * STELLARIS 8 ... CW and / or pulsed lasers, optional white light laser (WLL) laser (~440 - ~780nm, see data sheet) and AOBS, TauSense "fast FLIM" (fluorescence lifetime imaging microscopy) AND FALCON "fluorescence lifetime contrast" (fast FLIM, Phasors, other fancy FLIM stuff). All three models can have X1 port with additional detectors, such as two S dtectors or four APDs (avalanche photodiodes, which have ~80% quantum efficiency in red-NIR but slower photon counting and dynamic range per detector than S detector). GM thinking: we would love to have a STELLARIS 8 confocal microscope here. Potentially an NIH shared instrumentation grant proposal (May 2022?). If our user base does not need all the "8's" capabilities, a "5" or a "as demo'd" model would still enable much more productivity than our workhorse Leica SP8 confocal microscope (that was purchased within budget constraints and sensible choice for functionality and limiting annual service contract cost). If you or anyone you know have a bunch of money to donate for us to buy a STELLARIS, that would be awesome (we would trade in the SP8 and uses the same space). |
**
January 31, 2019: Ross S910 room hosting the ACCM Leica SP8 confocal microscope has been renovated with (i) blackout curtains for each microscope station, (ii) three pairs of track lights for each station, railings for the air ducts for HEPA filters.
|
Summer 2018 Leica Workshops hosted by Ross Fluorescence Image Core and ACCM Confocal Microscope Core * FALCON = FAst Lifetime CONtrast (Fluorescence Lifetime on HyD photon counting detectors) ==> potential future upgrade of SP8). Leica LAS X Navigator and TCS SP8 FALCON APPLICATION TALKS - June 19, 2018, 12:00-1:15 PM NAVIGATOR - HANDS ON DEMONSTRATION TIME - June 19-20 FALCON - Seminar, Geoff Daniels, Monday August 13. FALCON - HANDS ON DEMONSTRATION TIME - August 13-16 (Mon - Wed)
For Confocal applications, call or email Geoff Daniels (senior sales): Dvir Blivis (new regional confocal salesperson). Dvir.Blivis@leica-microsystems.com For Widefield applications, call or email Paula Cranfill: Regional leica team members include: Sarah Crowe - confocal applications Jessica Calafati - regional manager. Several field service engineers. *** George's Conclusions: * The new 3rd generation HyD detectors count faster than our SP8's "2nd gen" HyD's ... we would love if someone (SOM, DoM, philanthropy, anyone is welcome ... I note we would need a lot of user hours to pay for both annual service contract and buy 3rd gen HyDs from the daytime hourly rate of $27/hr) could provide money to (i) buy one or more of the 3rd gen HyDs. the Leica SP8 scanhead can use four HyDs,so ideally buy FOUR new 3rd gen HyDs and hopefully get Leica to accept the two current 2nd gen in trade-in). * FALCON concept is interesting ... Fast FLIM data implies -- to me -- a big "data deluge" AND requires at least one pulsed laser (yes, we would love for someone to donate money for one or more pulsed lasers, also new, more powerful computer to deal with the data). * I propose "Fast Photon Counting" (FPC) for now: 3rd gen HyDs, used with our current lasers, get data acquired a lot more quickly. The "photon counting speed advantage" of 3rd gen HyD's vs current 2nd gen is non-trivial to figure out under real use conditions. I estimate 10x advantage for each 3rd gen HyD at less than 2x the cost of each 2nd gen HyD. THis should not be seen as 10x / 2x = 5x advantage, but rather: one time investment ---> 7+ years of user's benefiting from 10x greater throughput (I estimate lifetime of SP8 is 8+ years,could be even longer). |
|
20200709 Update: I described Leica's 3rd generation HyD's above. These are (estimate) 40% Quantum Efficiency (QE) and were introduced Spring 2018. We hosted SP8 FALCON demo August 2018. In May 2020 leica introduced new STELLARIS Confocal microscope (I think of it as 'SP11' since prior numbering was SP2, SP5, SP8). The new confocal microscope can use the 3rd gen HyD, or an NIR version (i.e. GaAs faceplate), and features a new detector called POWER HyD(TM). This new detector is a new design (ask your local Leica rep what is it), with the HyD(TM) now being just a trademark. That is POWER HyD(TM) is NOT a PMT-APD hybrid detector, but something else. The peak "PQE" ~ 55% (QE accountign for the fill factor of the detector), so higher than 3rd gen HyD (the 'raw' QE would be higher). A cool feature of STELLARIS confocal scan head is it can use FIVE (!!!!!) POWER HyD detectors internally; My understanding is the X1 port can currently take 2 POWER HyD(TM)s, for total of 7 ... I strongly urge Leica to introduce "X4" with four POWER HyD(TM) for total of 9 (5 internal, 4 external), or even X8 for total of 13. This would be an amazing microscope! POWER HyD(TM) are as fast or faster than 3rd gen HyD, so in fast FLIM mode, a whole lotta data (which 2020 computers can deal with: E-ATX motherboard, PCIE gen4, NVidia Ampere GPUs, 1+ Terabyte fast RAM, lots of PCIe gen 4 NVMe SSD drives (re ASUS Hyper M2 or GloTrends cards), 10Gbe Ethernet (maybe 40Gbe) to local 'distributed computing network'. With White light laser ("WLL", Leica typically offers 440-900 nm excitation) and NKT Photonics "EXTEND-UV" to get down to 350nm excitation, STELLARIS/WLL/EXTEND-UV/POWER5internal/POWERX4/fast computing infrastructure, would be amazing. I also note that STELLARIS is STED capable (and STED benefits from flast FLIM, fast photon counting). |
|
20200902 Update ... information on our recommendation to users for using Photon Counting on our Leica SP8, and how settigns impact data: Photon counts: You control the amount of signal by: How much fluorescent dye (or fluorescent protein) is present in each region of your specimen.
I do note the HyD detectors have some background “counts” even with no laser power … this is low and random (i.e. 10 line accumulation most pixels will be zero, a few will be 1, very few will be 2), so can mostly ignore. Upshot: you can get a “good looking” image, even with just a few photons (ex. 10); better image quality with more photons (ex. 100 to 300 … 300 needs to use 16-bit mode [12-bit would work, but the image file is going to be 16-bit inside LAS X and export to raw TIFF, so might as well use 16-bit format). If your photon counts of bright areas are over 1000, you could probably decrease line accumulation (and frame accumulation).
The HyVolution2 computation (www.svi.nl Huygens deconvolution software with HyD detectors) always generates 16-bit image range (0 … ~65535), that is, scaled to the full range (to 65535) even if the photon counts are low (I usually do not use the one PMT on our SP8).
Trick for SP8 (eventually hopefully Leica will publish a ‘user generated application note’ I wrote about this): You can double the acquisition rate by using both HyD’s across the spectrum of one fluorophore, and then ADD the two channels in LAS X (or ImageJ or MetaMorph, etc). On our SP8, with two HyD detectors and one PMT in the middle, this could be HyD1 500-520nm, PMT 520-525nm (the minimum spectral band), HyD3 525-545nm (I would usually leave PMT off). Then add later (which is tedious in LAS X). This trick will be much more interesting with Leica STELLARIS 5 and 8 (launched May 2020: we would welcome upgrade!) where five of the new POWER HyD™ detectors could be used together (i.e. 500-540nm with adjacent 8nm bands; no need to waste any of the five internal detector positions with a PMT like our SP8 /// I also note that the STELLARIS external X1 port could be configured to be used with POWER detectors, would be great to have say 4 more POWER detectors externally for nine total [or even more with more money). Both “5” and “8” are ‘fast FLIM’ (fluorescence lifetime imaging microscopy) capable, “5” with “TauSense” modes (TauSeparation looks most useful to me; I note "5" does not provide raw data dump, so biophysicists may be bummed out if only have access to "5", but most biologists do not need the massive amount of raw data deluge of "8', and the money could be spent on more detectors for the "5"), “8” with "full data deluge" (~125 time intervals of ~100 picoseconds [0.1 ns] per 12.5 nanosecond pulse interval), TauSense and also FALCON (fluorescence lifetime contrast) and lifetime Phasor plots (‘set phasor to stun’! re: Star Trek). |
***
June 13, 2018: Leica SP8 is now in Ross S910A (S = Service corridor). Access is by scan card reader and SP8 is for fully tained users or users getting training from Dr. McNamara (or fee for service to have Dr. McNamara 'do' the imaging).
June 1, 2018: (see also above) Leica SP8 calendar/scheduler moved from Ross FIC (this site) to iLab Organizer as of June 1, 2018 (this scheduler booked all the time or calendar removed).***
We are also adding billing data per month per lab starting Feb 1, 2018 - May 31, 2018. Each P.I. will be billed a block of time each month (ex. Prof. Cyrus David Mintz, March 2018, 51 hours, in one time block).
January 5, 2018 news: ACCM's image core will be activated on iLab Organizer. Congratulations! We will continue to manage and host the microscope. The scheduling will move to iLab when the ACCM's iLab page is activated.
|
McNamara 20220106H 20210302U ACCM Leica SP8 Confocal Microscope Quick Tips on Starting Finishing User Session * we also have printouts on the Leica table (and nicer formatting). èonly fully trained users should touch the instrument and PC. If not used ‘in a while’ arrange in advance refresher training from George.
Web page: http://confocal.jhu.edu/current-equipment/leica-sp8-confocal-microscope iLab: https://johnshopkins.corefacilities.org/schedules/345585# (JHU login)
àNew user training notes (see also New User Training QA QC SOP doc): We normally train new users in two 2 hour sessions, ideally consecutive days. Each user should contact George McNamara (email gmcnamara@jhmi.edu) the fluorophores and specimens they plan to use. George will set up the training reservations (Agilent iLab and GM’s electronic calendar). We start out first training session with our Eosin tissue section slide and mostly 20x/0.75 NA dry objective lens. GM configures settings, explaining briefly the settings (and saves settings – a feature in HyVolution2 mode of Leica LAS X), acquires; User then uses microscope to find a new field of view, acquires. We then pivot to the user specimen(s). GM configures settings, evaluates crosstalk, adjusts any settings if needed, explaining to user, saves settings – which user can load for future sessions. User then uses microscope, and acquires. GM then shows how to acquire Z-series (if user project needs – most do).
Use JHU iLabs ACCM Confocal Microscope for scheduling in advance. Its billing is in ½ hour intervals, rounded up.
Starting SP8 sessions: Sign in first. … Failure to sign in = Trespassing.
During session: Leica SP8 PC fast drive is K:, organized as K:\ACCM Users\PI Lab, K:\GU Users\PI Lab, etc … save to your lab’s folder “early and often”. I recommend saving with your name, data, specimen number (ex: MaryEG 20210302 slide01). Most research lab PC’s struggle to deal with >1 Gigabyte (GB) data files, so I recommend keeping each LIF “container file” to under 1 GB – ex: save and close project file after each slide, potentially after each coverglass.
Finishing SP8 sessions:
All lasers to stand by
Notes: Keep LAS running.
Typical settings to get the most out of the Leica SP8 confocal microscope for pinhole 1.0 Airy Unit:
1.0 Airy unit
0.66 Airy Unit (0r 0.5 Airy unit) Objective lens XY Pixel Z-Step (up) HyVolution2
XY Pixel Z-Step (up) HyVolution2 20x/0.75 NA dry 120 nm 360 nm On, standard
Use the 63x lens 63x/1.4 NA oil (R.I. 1.518) 50 nm 150 nm On, standard resolution
40 nm 120 nm On, high resolution Optimization tips: http://confocal.jhu.edu/mctips/confocalsweetestspot (pinhole 1.0 and 0.66 Airy Units). |
|
|
Leica LAS NAVIGATOR -- a wizard for both tile scan & stitching and for multi-well plates -- can be used without selecting any of these 'templates'. We have a NAVIGATOR license. It is accessible from TCS SP8 mode only - the "many tiles" icon to the right of tile scan and mark&find ("NAV" is not visible in HyVolution2). When done with NAVIGATOR, please return to HyVolution2, turn off "red chain lock", and set speed to 600 Hz (from 700 Hz). Only 91.84 degrees (the rotation angle that matches the eyepiece and XY stage) and 1.94 degrees (which "NAV" will default to) work correctly. You can toggle back and forth between "NAV" and TCS SP8 modes (GUI = Graphical User Interface) - most LAS X users will find configuring scan settigns and spectral settigns simpler in the more familiar TCS SP8 "mode".
|

